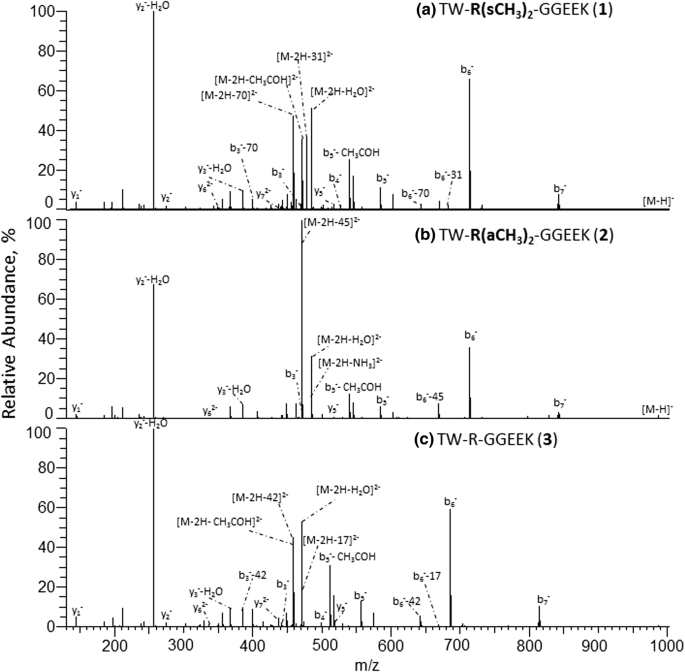
Negative Ion Mode Collision-Induced Dissociation for Analysis of Protein Arginine Methylation | SpringerLink
Model of tool function biotoolsSchema follows a simple model of tool... | Download Scientific Diagram

Extensive and Accurate Benchmarking of DIA Acquisition Methods and Software Tools Using a Complex Proteomic Standard | Journal of Proteome Research

Mixed-mode ion exchange-based integrated proteomics technology for fast and deep plasma proteome profiling - ScienceDirect
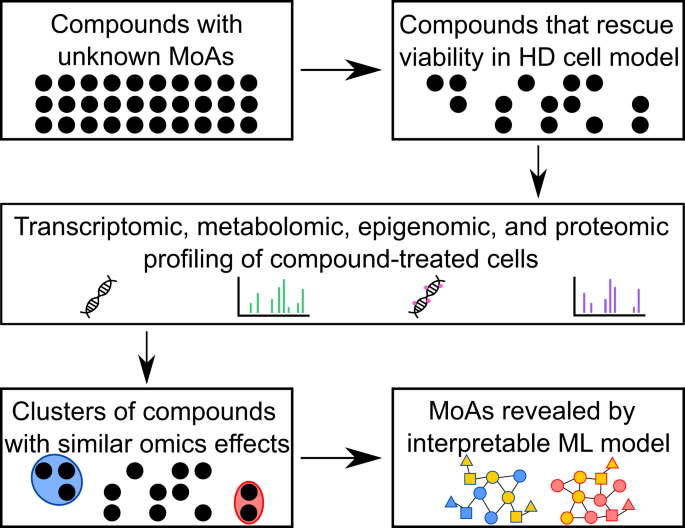
A Multi-Omics Interpretable Machine Learning Model Reveals Modes of Action of Small Molecules | Scientific Reports

A Simplified Flowchart for the Sequest Algorithm Showing the Process by... | Download Scientific Diagram
Differential Scanning Fluorometry Signatures as Indicators of Enzyme Inhibitor Mode of Action: Case Study of Glutathione S-Transferase | PLOS ONE

Differential procoagulatory response of microvascular, arterial and venous endothelial cells upon inflammation in vitro - Thrombosis Research

ProLuCID: An improved SEQUEST-like algorithm with enhanced sensitivity and specificity - ScienceDirect
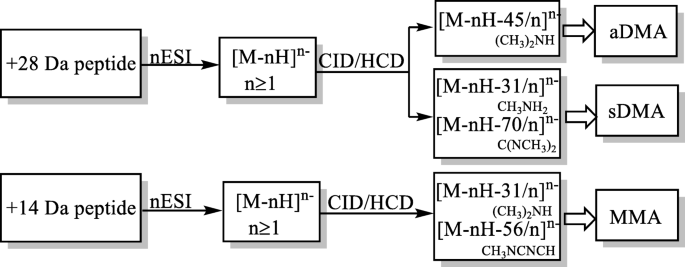
Negative Ion Mode Collision-Induced Dissociation for Analysis of Protein Arginine Methylation | SpringerLink

Analysis of protein digests by transmission-mode desorption electrospray ionization mass spectrometry with ultraviolet photodissociation - ScienceDirect

Widely accepted modes of Hfq activity.a | Hfq in association with a... | Download Scientific Diagram

Mascot and SEQUEST are complementary analysis tools for interpreting... | Download Scientific Diagram
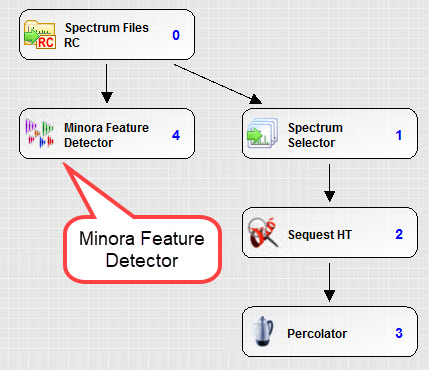
Configuring Proteome Discoverer 2.2 and forward for Compatibility with Scaffold – Proteome Software Technical Help Center